Visualization Results
Three Clusters Example
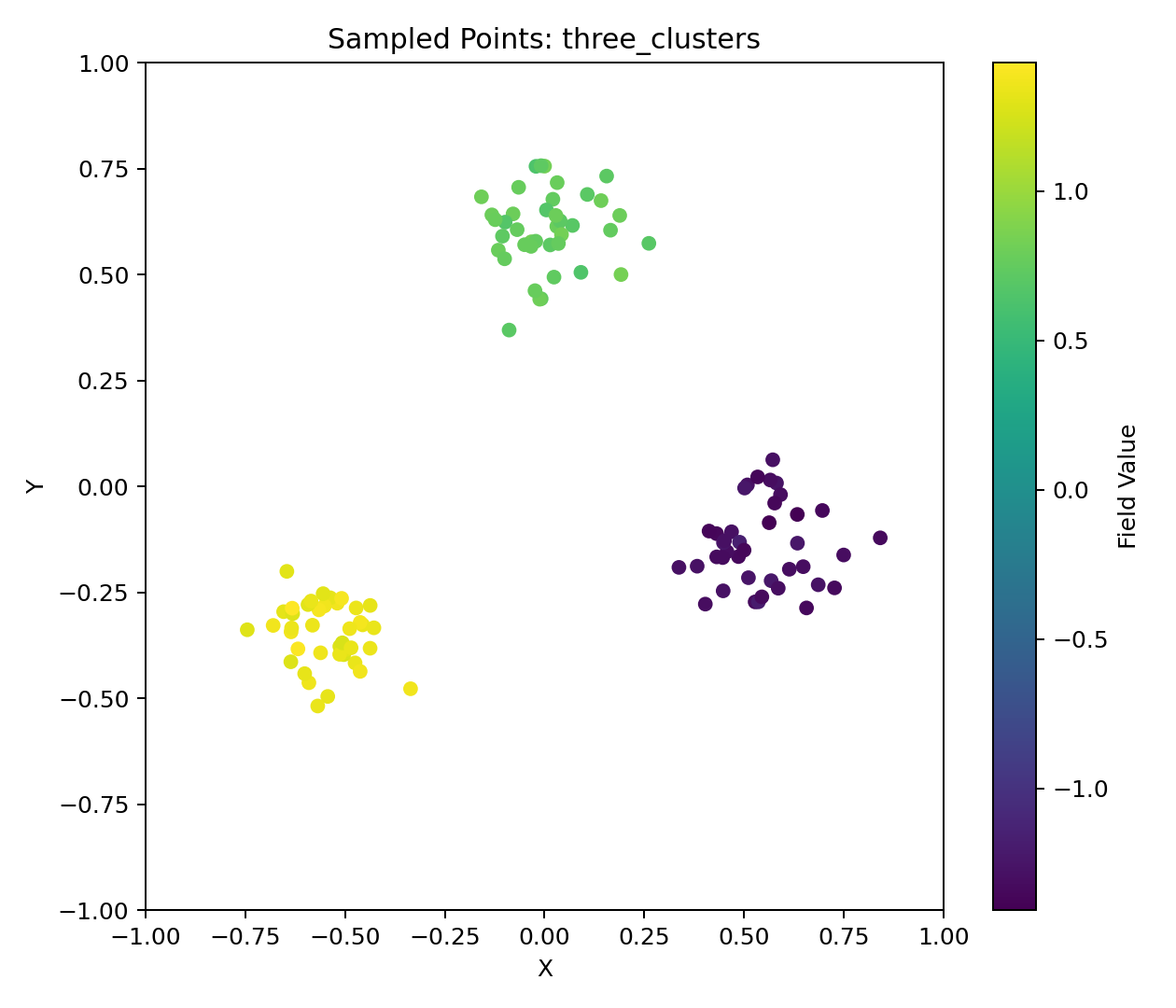
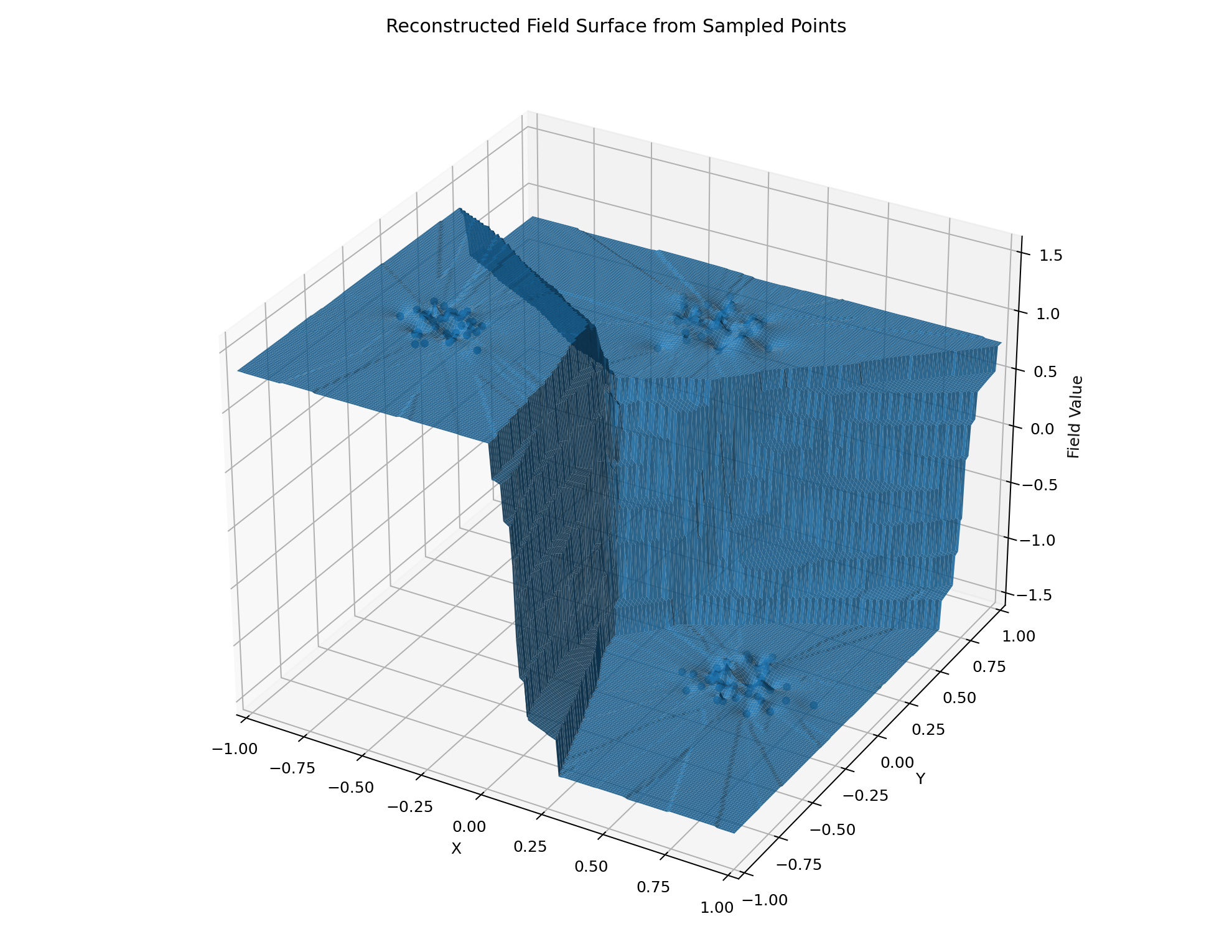
Zero Grid + Three Clusters Example
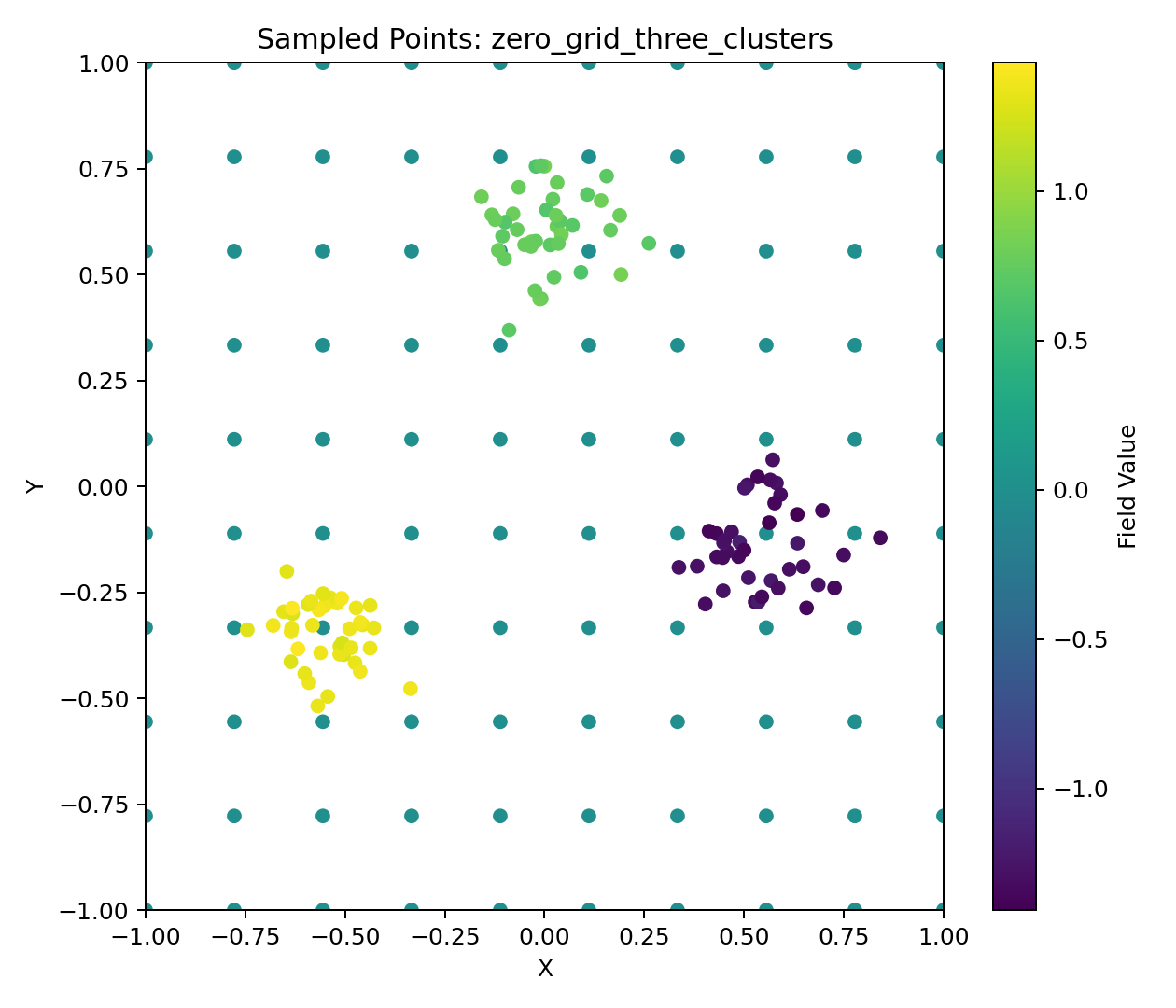
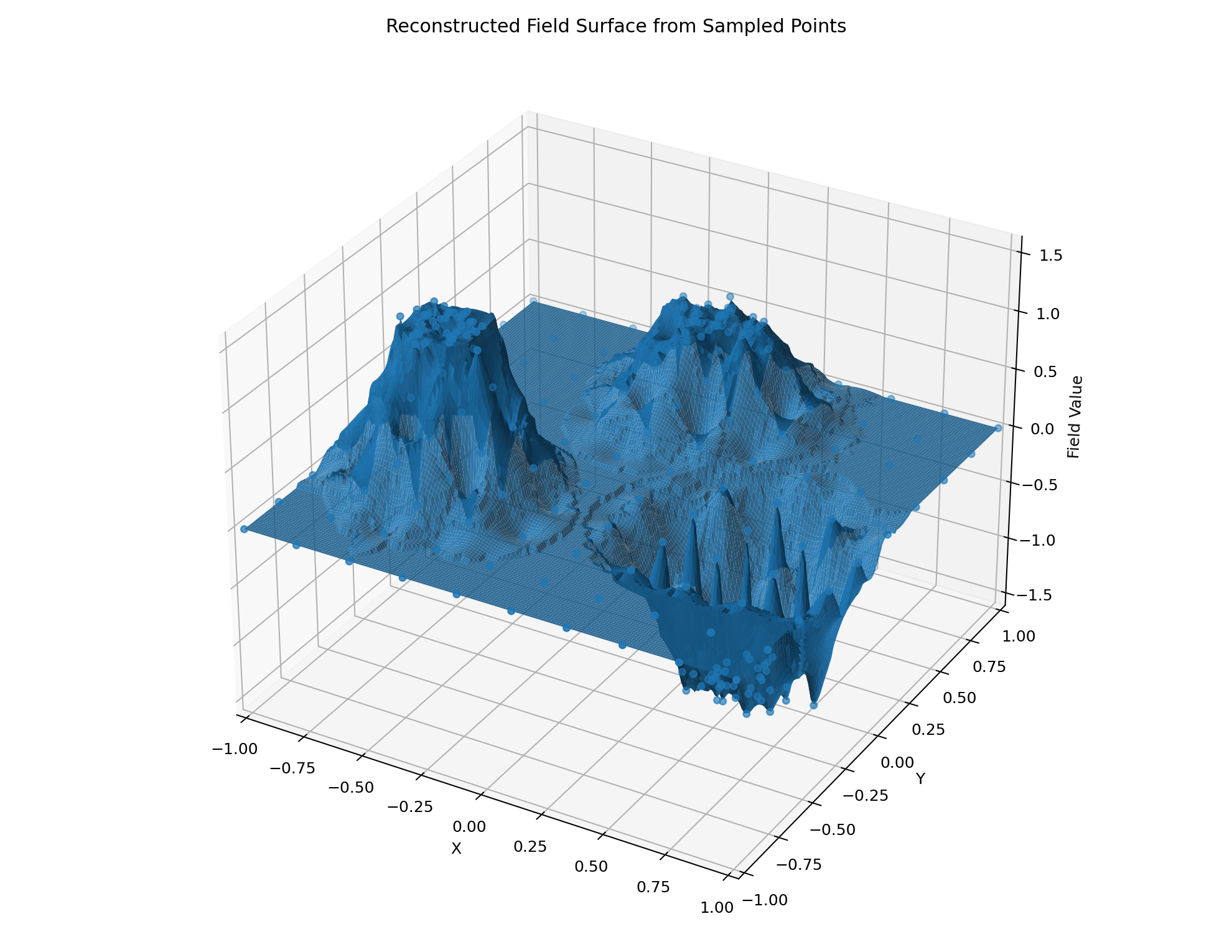
The goal of this project is to illustrate the reconstruction of a spatial field from scattered sample points. We focus on two example datasets:
This project shows how inverse distance weighting (IDW) can reconstruct a surface from scattered points.
The solution consists of Python scripts that:
We implemented modular functions to generate clusters, overlay grids, write CSVs, plot scatter points, and reconstruct surfaces.




The field reconstruction algorithm uses Inverse Distance Weighting (IDW) to interpolate scattered points onto a grid. Each grid point is estimated as a weighted average of nearby sampled points, with weights inversely proportional to distance raised to a power.
Steps:
This approach smoothly reconstructs the field while respecting local variations.
#!/usr/bin/env python3
from __future__ import annotations
import argparse
from pathlib import Path
import matplotlib
matplotlib.use('Agg')
import matplotlib.pyplot as plt
import numpy as np
from scipy.spatial import cKDTree
def load_points(path: str | Path):
data = np.genfromtxt(path, delimiter=',', names=True)
x = np.asarray(data['x'], dtype=float)
y = np.asarray(data['y'], dtype=float)
values = np.asarray(data['value'], dtype=float)
return x, y, values
def build_idw_interpolator(x, y, values, neighbors=8, power=2.0, eps=1e-12):
points = np.column_stack([x, y])
tree = cKDTree(points)
k = max(1, min(int(neighbors), len(points)))
def query(xq, yq):
q = np.column_stack([np.ravel(xq), np.ravel(yq)])
distances, indices = tree.query(q, k=k)
if k == 1:
result = values[np.asarray(indices)]
return result.reshape(np.shape(xq))
distances = np.asarray(distances, dtype=float)
indices = np.asarray(indices)
exact = distances[:, 0] < eps
safe_distances = np.where(distances < eps, eps, distances)
weights = 1.0 / np.power(safe_distances, power)
weighted = np.sum(weights * values[indices], axis=1) / np.sum(weights, axis=1)
if np.any(exact):
weighted[exact] = values[indices[exact, 0]]
return weighted.reshape(np.shape(xq))
return query
def make_plot(x, y, values, output_path, grid_size=180, neighbors=8, power=2.0):
interpolator = build_idw_interpolator(x, y, values, neighbors=neighbors, power=power)
limit = max(np.max(np.abs(x)), np.max(np.abs(y)), 1.0)
grid_x = np.linspace(-limit, limit, grid_size)
grid_y = np.linspace(-limit, limit, grid_size)
X, Y = np.meshgrid(grid_x, grid_y)
Z = interpolator(X, Y)
fig = plt.figure(figsize=(11, 8.5))
ax = fig.add_subplot(111, projection='3d')
ax.plot_surface(X, Y, Z, rstride=1, cstride=1, linewidth=0, antialiased=True, alpha=0.9)
ax.scatter(x, y, values, s=18, depthshade=True)
ax.set_title('Reconstructed Field Surface from Sampled Points')
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Field Value')
ax.set_xlim(-limit, limit)
ax.set_ylim(-limit, limit)
fig.tight_layout()
fig.savefig(output_path, dpi=180)
plt.close(fig)
def main():
parser = argparse.ArgumentParser(description='Visualize a scattered scalar field as a 3D surface.')
parser.add_argument('--input', required=True, help='CSV file with columns x,y,value')
parser.add_argument('--output', required=True, help='Output image path, e.g. field_surface.png')
parser.add_argument('--grid-size', type=int, default=180, help='Resolution of the rendered grid')
parser.add_argument('--neighbors', type=int, default=8, help='Number of nearest points used by IDW')
parser.add_argument('--power', type=float, default=2.0, help='IDW distance falloff power')
args = parser.parse_args()
x, y, values = load_points(args.input)
make_plot(x, y, values, args.output, grid_size=args.grid_size, neighbors=args.neighbors, power=args.power)
if __name__ == '__main__':
main()
The generate_examples.py orchestrates data creation, plotting, and calling the visualizer:
#!/usr/bin/env python3
from __future__ import annotations
import argparse
import csv
import subprocess
import sys
from pathlib import Path
import matplotlib
matplotlib.use('Agg')
import matplotlib.pyplot as plt
import numpy as np
APP_DIR = Path(__file__).parent
EXAMPLES_DIR = APP_DIR / 'examples'
VISUALIZER = APP_DIR / 'visualize_field.py'
def clear_examples_directory(path: Path) -> None:
path.mkdir(parents=True, exist_ok=True)
for item in path.iterdir():
if item.is_file() or item.is_symlink():
item.unlink()
def cluster_specs():
return [
{"center": (-0.55, -0.35), "value": 1.35, "n": 40, "spread": 0.10},
{"center": (0.55, -0.15), "value": -1.30, "n": 40, "spread": 0.10},
{"center": (0.00, 0.60), "value": 0.75, "n": 40, "spread": 0.09},
]
def generate_three_clusters(seed: int = 42):
rng = np.random.default_rng(seed)
xs = []
ys = []
zs = []
for spec in cluster_specs():
cx, cy = spec['center']
x = rng.normal(cx, spec['spread'], spec['n'])
y = rng.normal(cy, spec['spread'], spec['n'])
z = rng.normal(spec['value'], 0.05, spec['n'])
xs.append(np.clip(x, -1.0, 1.0))
ys.append(np.clip(y, -1.0, 1.0))
zs.append(z)
return np.concatenate(xs), np.concatenate(ys), np.concatenate(zs)
def generate_zero_grid_three_clusters(seed: int = 42):
x_clusters, y_clusters, z_clusters = generate_three_clusters(seed=seed)
base = np.linspace(-1.0, 1.0, 10)
X, Y = np.meshgrid(base, base)
x0 = X.ravel()
y0 = Y.ravel()
z0 = np.zeros_like(x0, dtype=float)
x = np.concatenate([x0, x_clusters])
y = np.concatenate([y0, y_clusters])
z = np.concatenate([z0, z_clusters])
return x, y, z
def write_csv(x, y, z, path: Path) -> None:
with path.open('w', newline='') as f:
writer = csv.writer(f)
writer.writerow(['x', 'y', 'value'])
writer.writerows(zip(x, y, z))
def make_scatter_plot(x, y, z, output_path: Path, title: str) -> None:
fig = plt.figure(figsize=(7, 6))
ax = fig.add_subplot(111)
scatter = ax.scatter(x, y, c=z, s=28)
ax.set_title(title)
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_xlim(-1.0, 1.0)
ax.set_ylim(-1.0, 1.0)
fig.colorbar(scatter, ax=ax, label='Field Value')
fig.tight_layout()
fig.savefig(output_path, dpi=180)
plt.close(fig)
def run_visualizer(csv_path: Path, output_path: Path, grid_size: int, neighbors: int, power: float) -> None:
subprocess.run(
[
sys.executable,
str(VISUALIZER),
'--input', str(csv_path),
'--output', str(output_path),
'--grid-size', str(grid_size),
'--neighbors', str(neighbors),
'--power', str(power),
],
check=True,
)
def render_example(name: str, x, y, z, grid_size: int, neighbors: int, power: float) -> None:
csv_path = EXAMPLES_DIR / f'{name}.csv'
scatter_path = EXAMPLES_DIR / f'{name}_scatter.png'
surface_path = EXAMPLES_DIR / f'{name}_surface.png'
write_csv(x, y, z, csv_path)
make_scatter_plot(x, y, z, scatter_path, title=f'Sampled Points: {name}')
run_visualizer(csv_path, surface_path, grid_size=grid_size, neighbors=neighbors, power=power)
print(f'Created {csv_path.name}, {scatter_path.name}, and {surface_path.name}')
def main() -> None:
parser = argparse.ArgumentParser(
description='Generate example datasets, scatter plots, and reconstructed surface images.'
)
parser.add_argument(
'--example',
choices=['all', 'three_clusters', 'zero_grid_three_clusters'],
default='all',
help='Which example set to generate',
)
parser.add_argument('--cluster-seed', type=int, default=42, help='Seed for the cluster-based examples')
parser.add_argument('--grid-size', type=int, default=180, help='Resolution of the rendered grid')
parser.add_argument('--neighbors', type=int, default=8, help='Number of nearest points used by IDW')
parser.add_argument('--power', type=float, default=2.0, help='IDW distance falloff power')
args = parser.parse_args()
clear_examples_directory(EXAMPLES_DIR)
if args.example in ('all', 'three_clusters'):
x, y, z = generate_three_clusters(seed=args.cluster_seed)
render_example(
name='three_clusters',
x=x,
y=y,
z=z,
grid_size=args.grid_size,
neighbors=args.neighbors,
power=args.power,
)
if args.example in ('all', 'zero_grid_three_clusters'):
x, y, z = generate_zero_grid_three_clusters(seed=args.cluster_seed)
render_example(
name='zero_grid_three_clusters',
x=x,
y=y,
z=z,
grid_size=args.grid_size,
neighbors=args.neighbors,
power=args.power,
)
print(f'Finished. Outputs are in: {EXAMPLES_DIR}')
if __name__ == '__main__':
main()
Download the complete project, including Python scripts, CSV files, visualizations, and this web page:
Download field_visualization_two_examples.zip